|
For the expansion of this macrovintertebrates.org we've needed to develop a robust content management system that scales to accommodate a more intricate data structure and authoring needs of a larger project team, as well as support new kinds of web interactions and usage tracking. Our new Content and Annotation Tool (CAT), allows the users to input the necessary data for each macroinvertebrate in our collection and locate custom multimedia annotations in a multiscalar image space. Organization was a priority in this redesign because we wanted to make sure that we could keep track of the complex nested hierarchies in taxonomy so that the various diagnostic characters at each level would be attached to each species, and comparable across species. Throughout our design process we received a lot of input and feedback from our entomologists to understand and align the system to the way they understand and organize their data. Some of the issues that we wanted to resolve in this redesign of the backend system include:
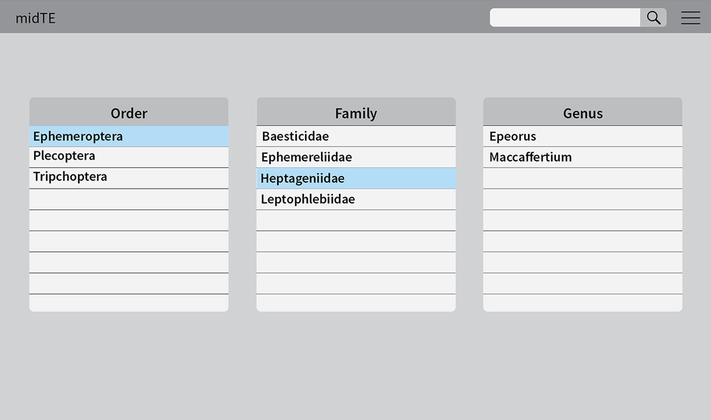
We started this process back in March by creating sketches and paper prototypes of potential organizational systems. We then moved onto building wireframes that were used to demonstrate how to input an entry into the system and annotate diagnostic characters on the Gigpan images. In building this clickable walkthrough, we were able to share this with Maddie, our entomologist at Clemson and receive feedback on how she was using the system. We also checked in with Chris, our developer at various points to get his feedback on the technical aspects of the system, such as how to manage multiple users and how to search for existing entries and information within the database. The following are some screenshots of the initial content management we prototyped. The landing page for the prototype was based on an iTunes style browser that enables entries to be viewed in a hierarchal order based on taxonomy. When an order, such as Ephemeroptera is selected from the box on the left hand, all the families that have already been entered in the system will displayed in the center box. When a family from within the center box has been selected, such as Heptageniidae, all the genera that have been entered under this family would be displayed. Another goal in redesigning the tool was to improve consistency when entering information in the backend system. We addressed this issue by creating tabs that covered “Panel Information”, “Diagnostic Characters”, and “IP Tool”. Panel Information contains all the entry fields necessary for the overlay panel at each level which includes general description and related media. Diagnostic characters is a list of features that are added to each entry specific to its taxonomic hierarchy. For example, at the order level for Ephemeroptera, the feature, “Three Tails” is added that will ripple through the system anytime any subsequent entry is added at the family or genus level for Ephemeroptera. In these tabs, there is also a flag next to each data entry section that can be turned on or off to signify that the content needs to be returned to, whether to be looked over or to finish. This tool can be used by multiple users to communicate points in the data entry system that needs to be revisited before publishing. In the IP Tool tab, these diagnostic characters can then be annotated and modified based on specific species. In this redesign, we wanted to design a conceptual framework and data structure our developer could reference. The tool that Chris developed for the backend management system incorporates aspects of our previous redesign. To kickoff the CAT, Chris our developer did a scan of existing open source CMS systems to establish whether this project required a custom build or could be developed upon an existing framework.. Various platforms were considered, such as Hydra, but Collective Access (CA) was selected based on its ability to be customized to suit the specific needs and requirements of this project. CA is an open source software for management of collections in museum and archival settings. One of the main advantages of using an existing software to build our system on was that the tool did not have to be built from scratch. Instead, CA already has developed features such as multiple authors, versioning, user interfaces, data input validations, integration with maps, and media uploads. Although there are some limitations in CA due to it being a platform not specifically designed for our purposes, Chris was able to iterate on the schema based on feedback from the scientists and the designers to improve on the tool. Walkthroughs have been conducted with our entomologist to verify that the data entry process addresses all the necessary information that needs to be entered at each level. Through these walkthroughs, Chris has made adjustment such as reorganizing the UI and enlarging a button to add a feature, along with addressing relationships between diagnostic characters. A staging website was also created so that users adding content to the backend are able to preview what is on the website without having to directly publish the material. At the current moment, iterations are still being made to the site, but overall our system is able to incorporates aspects of taxonomic hierarchies within a content management tool. This CAT screenshot shows the form for creating a new diagnostic character at the family level. Once it has been entered here, it can be added as a diagnostic character for families that share the same character. It is similar in concept to our diagnostic character tab from our prototype, but uses the built in interface for CA. This allows us to make iterations to the overall design and process with the scientists, developers and designers rather than spending extra time to build the system from scratch.
Comments are closed.
|
Project TeamAn interdisciplinary team Categories
All
Archives
June 2023
|

 RSS Feed
RSS Feed
